Segmenting the picture of a raccoon face in regions¶
This example uses Spectral clustering on a graph created from voxel-to-voxel difference on an image to break this image into multiple partly-homogeneous regions.
This procedure (spectral clustering on an image) is an efficient approximate solution for finding normalized graph cuts.
There are two options to assign labels:
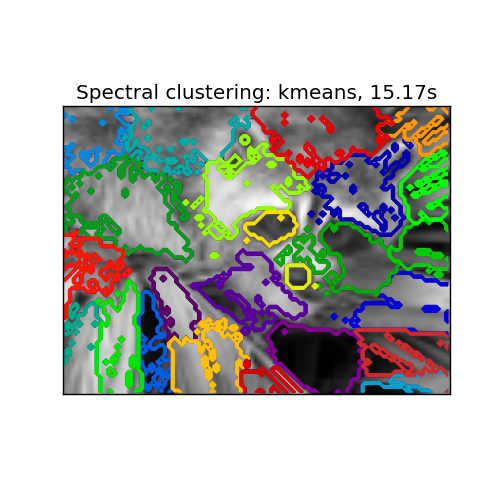
- with ‘kmeans’ spectral clustering will cluster samples in the embedding space using a kmeans algorithm
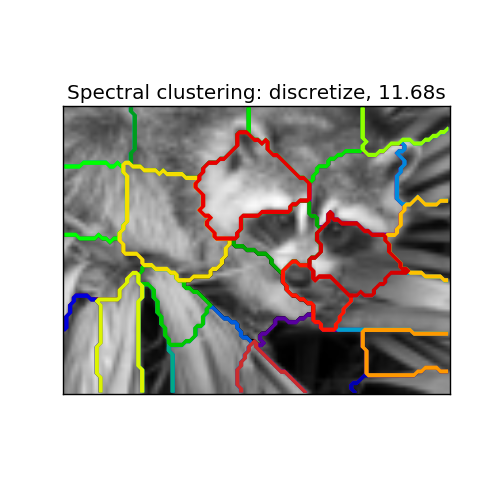
- whereas ‘discrete’ will iteratively search for the closest partition space to the embedding space.
Script output:
Spectral clustering: kmeans, 15.17s
Spectral clustering: discretize, 11.68s
Python source code: plot_face_segmentation.py
print(__doc__)
# Author: Gael Varoquaux <gael.varoquaux@normalesup.org>, Brian Cheung
# License: BSD 3 clause
import time
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
from sklearn.feature_extraction import image
from sklearn.cluster import spectral_clustering
from sklearn.utils.testing import SkipTest
from sklearn.utils.fixes import sp_version
if sp_version < (0, 12):
raise SkipTest("Skipping because SciPy version earlier than 0.12.0 and "
"thus does not include the scipy.misc.face() image.")
# load the raccoon face as a numpy array
try:
face = sp.face(gray=True)
except AttributeError:
# Newer versions of scipy have face in misc
from scipy import misc
face = misc.face(gray=True)
# Resize it to 10% of the original size to speed up the processing
face = sp.misc.imresize(face, 0.10) / 255.
# Convert the image into a graph with the value of the gradient on the
# edges.
graph = image.img_to_graph(face)
# Take a decreasing function of the gradient: an exponential
# The smaller beta is, the more independent the segmentation is of the
# actual image. For beta=1, the segmentation is close to a voronoi
beta = 5
eps = 1e-6
graph.data = np.exp(-beta * graph.data / graph.data.std()) + eps
# Apply spectral clustering (this step goes much faster if you have pyamg
# installed)
N_REGIONS = 25
#############################################################################
# Visualize the resulting regions
for assign_labels in ('kmeans', 'discretize'):
t0 = time.time()
labels = spectral_clustering(graph, n_clusters=N_REGIONS,
assign_labels=assign_labels, random_state=1)
t1 = time.time()
labels = labels.reshape(face.shape)
plt.figure(figsize=(5, 5))
plt.imshow(face, cmap=plt.cm.gray)
for l in range(N_REGIONS):
plt.contour(labels == l, contours=1,
colors=[plt.cm.spectral(l / float(N_REGIONS))])
plt.xticks(())
plt.yticks(())
title = 'Spectral clustering: %s, %.2fs' % (assign_labels, (t1 - t0))
print(title)
plt.title(title)
plt.show()
Total running time of the example: 29.08 seconds ( 0 minutes 29.08 seconds)